
Faculty Recruiting Seminar
Monday, December 17, 3:30-4:30pm, WEL 2.122
Postdoctoral Researcher
UC San Francisco
PhD 2013, UC Davis
Cellular processes are orchestrated by a large number of biomolecules in a spatially and temporally coordinated manner within a tiny volume. To uncover the underlying organizational principles and their functional relevance, we take microscopy visualization as the primary approach to systematically map the spatial localization, temporal dynamics and activity profiles of proteins and nucleic acids.
h-index: 19 Total Citations: 223 (Google Scholar Citations, Nov. 2018)
Faculty Recruiting Seminar
Tuesday, December 11, 3:30-4:30pm, NHB 1.720
Postdoctoral Research Fellow
Stanford University
PhD 2014, Boston University
The Moerner Laboratory utilizes laser spectroscopy and microscopy of single molecules to probe biological processes, one biomolecule at a time. Primary thrusts include development and application of fluorescence microscopy far beyond the optical diffraction limit by PALM/STORM and STED approaches, invention and validation of methods for precise and accurate 3D optical microscopy in cells, and trapping of single biomolecules in solution for extended study. These approaches are applied to explore protein localization patterns in bacteria, to measure structures of amyloid aggregates in cells, to define the behavior of signaling proteins in the primary cilium, to quantify photodynamics for photosynthetic proteins and enzymes, and to observe the dynamics of DNA and RNA in cells and viruses.
h-index: 6 Total Citations: 242 (Google Scholar Citations, Nov. 2018)
h-index: 4 Total Articles: 7 Total Citations: 89 (Web of Science, Nov. 2018)
Centennial Lecture
Thursday, Nov. 29, 3:30 - 4:30pm, NHB 1.720 
Distinguished Professor of Chemistry
University of Utah
The White group is an interdisciplinary team that applies principles grounded in electrochemistry to advance understanding of a wide variety of physical systems. Systems currently investigated within the group span the physical, analytical and biological sciences. These include nano-pore/pipette based sensing, measuring and delivery of nanoscale particles; protein ion channel electrical measurements of nucleic acids; theoretical and electrochemical scanned probe investigations of solid-state batteries; experimental measurements and theoretical analysis of ionic transport in confined geometries; and fundamental studies of electrochemically generated nanobubbles.
Publications (Google Scholar Citations)
h-index: 70 Total Articles: 386 Total Citations: 18,655 (Web of Science, Nov. 2018)
h-index: 73 Total Citations: 17,444 (Google Scholar Citations, Nov. 2018)
Thursday, Nov. 8, 3:30pm - 5:00pm, NHB 1.720 
Assistant Professor of Chemistry
Ohio State University
Our research focuses on the critical role that surface electron dynamics and interfacial charge transfer have on the selectivity and efficiency of catalytic energy conversion processes. Toward this goal my group has developed femtosecond soft x-ray reflection-absorption spectroscopy as a surface sensitive probe of ultrafast electron dynamics. We also study the closely related chemical reaction kinetics on catalytic surfaces.
ORCID: https://orcid.org/0000-0001-6740-864X
h-index: 14 Total Articles: 25 Total Citations: 590 (Web of Science, Oct. 2018)
Friday, Nov. 2, 3:30pm - 5:00pm, WEL 2.122 
Professor of Chemistry
Kansas State University
Two of the major research aims in the Aikens group involve developing a theoretical understanding of the relationship between structure and properties of nanomaterials. These research areas include the structure and spectroscopic properties of gold and silver nanoparticles and nanoparticle arrays and the reactivity of nanostructured metal oxide particles.
h-index: 29 Total Articles: 88 Total Citations: 4617 (Web of Science, Oct. 2018)
Thursday, Nov. 1, 3:30pm - 4:30pm, NHB 1.720
Professor of Biochemistry
Vanderbilt University School of Medicine
The research interests of this laboratory are aimed at the investigation of biological processes involving the synthesis, modification, storage and degradation of certain peptides and proteins using modern mass spectrometric methods of analysis to follow molecular events. In recent years there has been a great amount of interest in investigating the biochemical events involved in the metabolism of peptides, primarily in the brain and gut of mammals, encompassing the enzymatic breakdown of these peptides, their production from peptide and protein precursors, and the disruption of these processes by certain xenobiotics. Modern mass spectrometric techniques are used in these studies, including electrospray and matrix-assisted laser desorption ionization mass spectrometry.
h-index: 75 Total Publications: 310 Total Citations: 18,622 (Web of Science, Oct. 2018)
Thursday, Oct. 25, 3:30pm - 4:30pm, NHB 1.720
Associate Professor of Chemistry
UC Riverside
Our research focuses on computational materials chemistry and nanoscience, with a long-term goal to achieve knowledge-based design of functional materials for a sustainable society. Current research topics:
ORCID: http://orcid.org/0000-0001-5167-0731
h-index: 63 Total Citations: 11,963 (Google Scholar Citations, Sep 2018)
Thursday, Oct. 11, 3:30pm - 4:30pm, NHB 1.720 
M.G. Mellon Distinguished Professor of Chemistry
Purdue University
The Zwier group focuses on problems at the interface between the traditional fields of molecular spectroscopy and dynamics. We develop and apply laser-based methods to study neutral molecules, ions, radicals, and molecular clusters of a size and complexity that test the large-molecule limits of current theories of chemical structure and dynamics.
Our current research interests have applications in the following three areas:
1. The investigation of conformational preferences, isomerization dynamics, and hydrogen bonding in biologically relevant neutral molecules, molecular ions, and molecular clusters. A particular current interest is synthetic foldamers, molecules designed to fold in well-defined ways that either mimic or complement the folding propensities of α-peptides found in nature.
2. The study of the photo-induced and discharge-driven chemistry and spectroscopy of molecules of importance in planetary atmospheres, particularly on Titan, one of the moons of Saturn.
3. The study of the chemistry, spectroscopy, and isomerization of molecules and radicals of importance in combustion, seeking to understand reactions and energy flow in aromatic derivatives that are intermediates along the pathway towards soot formation.
ORCID: https://orcid.org/0000-0002-4468-5748
h-index: 47 Total Publications: 197 Total Citations: 8668 (Web of Science, Sep. 2018)
Thursday, Sept 27, 3:30pm - 4:30pm, NHB 1.720 
Assistant Professor of Chemistry
MIT
Research in our group is inherently multidisciplinary; we combine tools from chemistry, optics, biology, and microscopy to develop new approaches to probe dynamics. We study dynamics in two classes of systems: biological and bio-inspired light-harvesting systems that are of interest to solar energy research and biomass production; and bacterial and mammalian receptor proteins that are targets for human therapeutics. To explore these systems, we use ultrafast transient absorption spectroscopy, single-molecule fluorescence spectroscopy, and develop model membrane systems.
h-index: 13 Total Publications: 20 Total Citations: 1025 (Web of Science, Aug. 2018)
Thursday, Sept. 20, 4:00-5:00pm, WEL 2.122 
Merck
Elizabeth Pierson currently works at the Research Laboratories, Merck. Elizabeth does research in Analytical Chemistry, Virology and Molecular Biology. Their current project is 'Analytical Research and Development of Formulated Small Molecule and Biological Pharmaceuticals.
Friday, April 20, 1:30pm - 2:30pm, NHB 1.720
Professor of Chemistry
Penn State University
The laboratory of Biointerfaces, led by Prof. Paul Cremer, is a multidisciplinary research team that works at the crossroads of biological interfaces, manomaterials, spectroscopy, and microfluidics. Biophysical and analytical studies are tied together through the employment of novel lab-on-a-chip platforms which enable high throughput/low sample volume analysis to be performed with unprecedented signal-to-noise. From neurodegenerative diseases to artificial hip implants, a huge variety of processes occur at biological interfaces. Our laboratory uses a wide variety of surface specific spectroscopies and microfluidic technologies to probe mechanisms of disease, build new biosensors against pathogens, and understand the molecular-level details of the water layer hugging a cell membrane.
h-index: 59 Total Publications: 134 Total Citations: 12,064 (Web of Science, Mar. 2018)
Thursday, April 5, 3:30pm - 4:30pm, WEL 2.122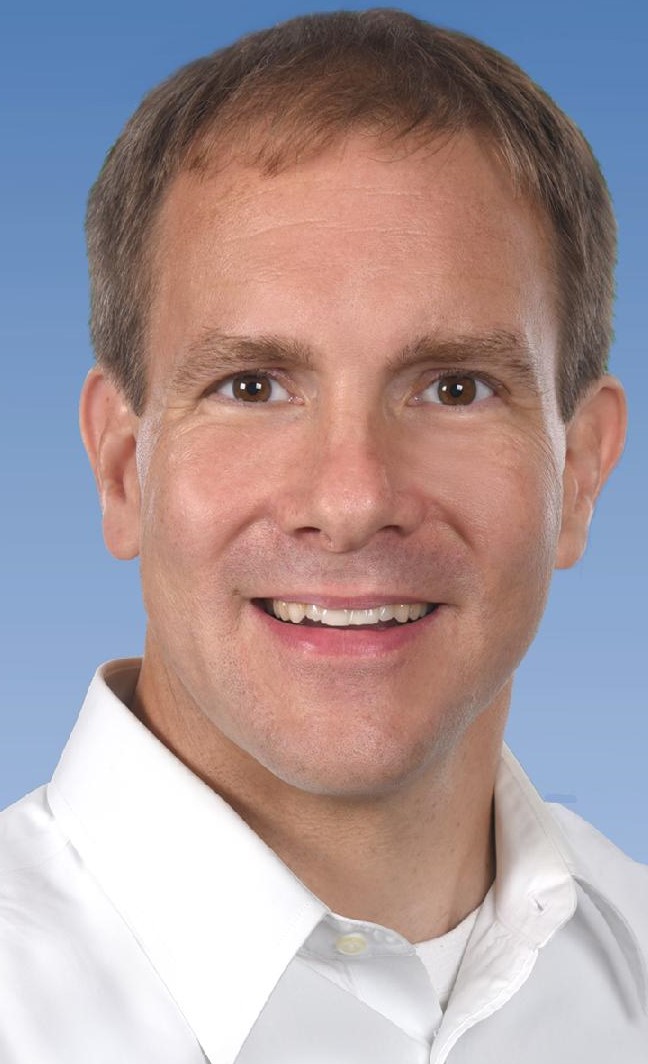 
Assistant Professor of Chemistry
UC Irvine
The central theme of the Ardo Group's research program is to understand and control reaction mechanisms at interfaces, with the goal of optimizing energy conversion for practical applications, including solar seawater desalination, solar fuels devices, photovoltaics, redox flow batteries, and fuel cells.
h-index: 20 Total Publications: 38 Total Citations: 2255 (Web of Science, Mar. 2018)
Thursday, March 29, 3:30pm - 5:00pm, WEL 2.122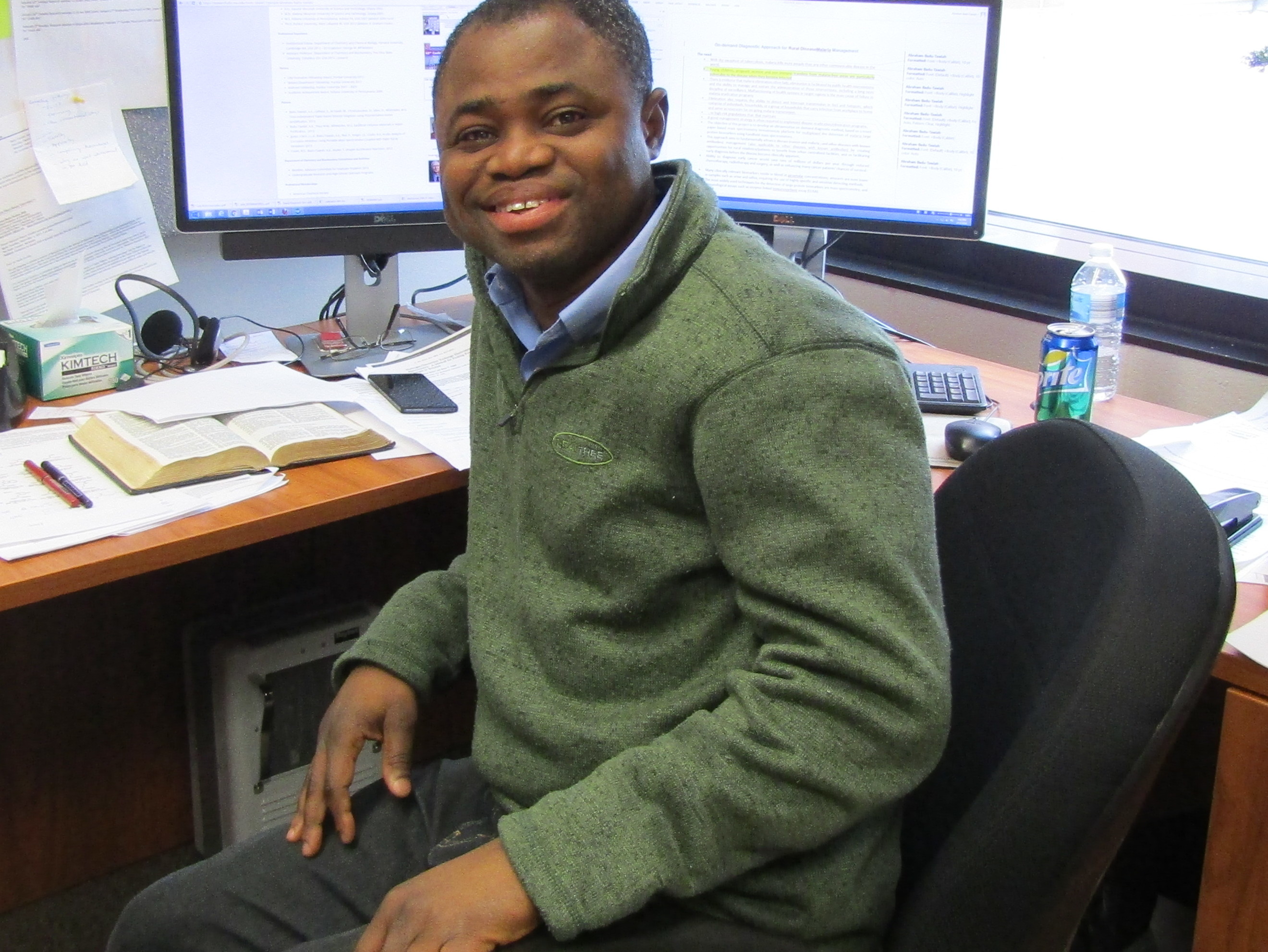 
Assistant Professor of Chemistry
Ohio State University
Recent innovation in mass spectrometry (MS) is the ability to generate intact molecular ions, focus and use them as ordinary reagents for organic reactions at ambient surface, outside the mass spectrometer. Most projects in the Badu lab build on this innovation, but instead of simple organic compounds, we focus on drugs and biomolecules of specific biological importance.
ORCID: https://orcid.org/0000-0001-8642-3431
h-index: 14 Total Publications: 25 Total Citations: 482 (Web of Science, Mar. 2018)
h-index: 14 Total Publications: 26 Total Citations: 538 (Scopus, Mar. 2018)
George and Pauline Watt Centennial Lectureship 
Monday, March 26, 2018, 03:30pm - 04:30pm, WEL 2.122
Distinguished Professor, Chemistry
UCLA
Physical/chemical: fluorescence correlation spectroscopy (FCS); high resolution cryo-electron microscopy (cryo-EM); magnetic and optical tweezers (Professors James Bowie (UCLA) and Doug Smith (UCSD)) maleimide and “click” chemistries for conjugating ligands to capsid protein; electrophoretic, sedimentation, and chromatographic separations and analyses of fluorescently-labeled RNA, protein, and RNA-protein complexes; labeling of RNA ends, and of capsid proteins, by <2nm gold particles; statistical-mechanical modeling.
Biological: agrobacterial transformation of plants for high-level expression of wildtype and mutant CCMV capsid protein; genetic engineering of RNA-virus-derived replicons from mammalian, insect, and plant viruses; transfection and infection of cultured cells for assays of RNA replication and protein expression levels; in vitro transcription and genetic engineering of viral genes and their protein products; cell-free translation of RNA, of virus-like particles, and of viruses.
Festschrift - J. Phys. Chem. B. (2016)
h-index: 69 Total Publications: 221 Total Citations: 20,814 (Web of Science, Mar. 2018)
h-index: 58 Total Publications: 178 Total Citations: 10,742 (Scopus, Mar. 2018)
Friday, March 23, 3:30pm - 5:00pm, WEL 2.122 
Faculty, Dept. of Chemistry
University of Rome La Sapienza
h-index: 25 Total Publications: 107 Total Citations: 1790 (Web of Science, Feb. 2018)
h-index: 26 Total Publications: 112 Total Citations: 1785 (Scopus, Feb. 2018)
h-index: 28 Total Citations: 2083 (Google Scholar Citations, Feb. 2018)
W. Albert Noyes, Jr. Distinguished Visiting Lectureship 
Thursday, March 1, 3:30pm - 4:30pm, WEL 2.122
Professor of Biochemistry, Biophysics and Structural Biology
UC Berkeley
The long-term goal of our research is to understand the structural and dynamic information encoded in the linear sequence of amino acids. Proteins undergo an incredible transformation from one-dimensional sequence information into complex three-dimensional shapes that carry out intricate cellular functions. We still, however, don't have enough biophysical knowledge to translate this sequence information into functional insights. For instance, many proteins share the same native structure yet their cellular dynamics and function, in other words their energy landscapes, are different. Our laboratory uses a combination of biophysical, structural and computational techniques to understand these features.
h-index: 42 Total Publications: 97 Total Citations: 7040 (Web of Science, Feb. 2018)
h-index: 42 Total Publications: 110 Total Citations: 7048 (Scopus, Feb. 2018)
Thursday, February 22, 3:30pm - 4:30pm, WEL 2.122
Professor
Emory University
Research in the Dyer group spans a broad range of problems in biophysical and bioinorganic chemistry. Links to the three main focus areas are included below: mapping protein folding energy landscapes, exploring the role of protein dynamics in enzymatic catalysis, and developing photocatalysts for solar water splitting and hydrogen production. We have also undertaken a collaboration with the Salaita lab, funded by DARPA, to exploit coupled enzymatic reactions for induced mechanical work.
Two unifying themes of this work are exploring the role of dynamics in protein structure and function and the development and application of new laser and spectroscopic tools for the study of protein dynamics. Our work effectively cuts across traditional disciplines with an emphasis on using quantitative physical methods to address biological problems. For example, our study of fast events in protein folding integrates efforts in mechanical engineering (microfluidics for ultrafast mixing), physical chemistry (spectroscopy, fast kinetics, physical models), molecular biology (mutants, protein folding models) and theoretical chemistry (MD simulations of folding). We emphasize the use of spectroscopic techniques with high structural specificity and time resolution, such as isotope edited infrared spectroscopy to elucidate the functional dynamics of proteins on all relevant timescales.
h-index: 50 Total Citations: 6897 (Google Scholar, Feb. 2018)
h-index: 45 Total Publications: 131 Total Citations: 5637 (Web of Science, Feb. 2018)
Wednesday, February 21, 3:30pm - 4:30pm, WEL 2.122 
Professor
Cornell University
Our research focuses on developing and applying state-of-the-art single-molecule methods to characterize and understand the properties of nanoscale materials and biological systems. Compared with traditional ensemble measurements, the single molecule approach removes ensemble averaging, so that distributions and fluctuations of molecular properties can be characterized and transient intermediates identified. The single-molecule techniques we employ include single-molecule fluorescence imaging, single-molecule FRET, single-molecule tracking, super-resolution localization microscopy, and magnetic tweezers. Our research program provides students with scientific training spanning from sophisticated microscopy/spectroscopy techniques, rigorous data analyses to protein and genetic engineering using modern molecular biology techniques, as well as nanotechnology and nanomaterials.
h-index: 31 Total Publications: 57 Total Citations: 4069 (Google Scholar, Feb. 2018)
Thursday, February 15, 3:30pm - 5:00pm, WEL 2.122 
Professor, Chemistry
Northwestern University
The Mirkin Research Group focuses on developing methods for controlling the architecture of molecules and materials on the 1 – 100 nm length scale, understanding their fundamental properties, and utilizing such structures to develop novel tools that can be applied in the areas of chemical and biological sensing, gene regulation, immunomodulation, lithography, catalysis, optics, and energy generation, storage, and conversion.
The Mirkin Research Group has pioneered the use of nanoparticle-biomolecule conjugates as synthons in materials science and the development of many nanoparticle-based extra- and intracellular biodiagnostic and therapeutic tools.
h-index: 132 Total Publications: 607 Total Citations: 87,238 (Web of Science, Jan. 2018)
 Highly Cited: 46 Articles
Highly Cited: 46 Articles
h-index: 135 Total Publications: 705 Total Citations: 90,931 (Scopus, Feb. 2018)
Centennial Lecture 
Thursday, February 8, 3:30pm - 4:30pm, WEL 2.122
Professor, Chemical Physiology
Scripps Research Institute
Research in the Yates lab is focused on the development and application of mass spectrometry-based proteomics techniques to a wide range of biological questions. Our lab has been instrumental in the evolution of the field to its current status, having pioneered many of the landmark advances that form the basis for prevailing proteomics practices, including shotgun proteomics (McCormack, A. L.; Schieltz, D. M.; Goode, B.; Yang, S.; Barnes, G.; Drubin, D.; Yates, J. R., III. Anal. Chem. 1997, 69, 767−776), database searching (SEQUEST, Eng, J. K.; McCormack, A. L.; Yates, J. R., III. J. Am. Soc. Mass Spectrom. 1994, 5, 976−989), and Multidimensional Protein Identification Technology (MudPIT, Washburn, M. P.; Wolters, D.; Yates, J. R., III. Nat. Biotechnol. 2001, 19, 242−247). We continue the drive to increase the scope, sensitivity and throughput of proteomics technologies and their application to biological questions.
Our research encompasses the areas of bioinformatics and software development, methods development and biological applications. The integration of all the elements in the proteomics pipeline within one lab facilitates advances in all of them.
The Yates lab has published more than 700 peer reviewed papers. Recent highlights include comprehensive proteomics studies revealing molecular mechanisms implicated in Cystic Fibrosis as well as identification of proteins capable of restoring function to mutated proteins in the disease, and investigations into affective disorders of the brain, including schizophrenia and depression.
ORCID: https://orcid.org/0000-0001-5267-1672
h-index: 132 Total Publications: 708 Total Citations: 69,178 (Web of Science, Jan. 2018)
Thursday, February 1, 3:30pm - 5:00pm, WEL 2.122 
Assistant Professor, Chemistry
MIT
h-index: 12 Total Publications: 27 Total Citations: 975 (Web of Science, Jan. 2018)
Thursday, January 25, 3:30pm - 4:30pm, WEL 2.122 
Associate Professor, Chemistry
University of Louisville
h-index: 35 Total Publications: 81 Total Citations: 3313 (Web of Science, Dec. 2017)
 Highly Cited Paper: Mieszawska, AJ et al. The synthesis and fabrication of one-dimensional nanoscale heterojunctions. Small, 3(5), 2007, 722-756. DOI: 10.1002/smll.200600727
Highly Cited Paper: Mieszawska, AJ et al. The synthesis and fabrication of one-dimensional nanoscale heterojunctions. Small, 3(5), 2007, 722-756. DOI: 10.1002/smll.200600727
Faculty Recruiting Seminar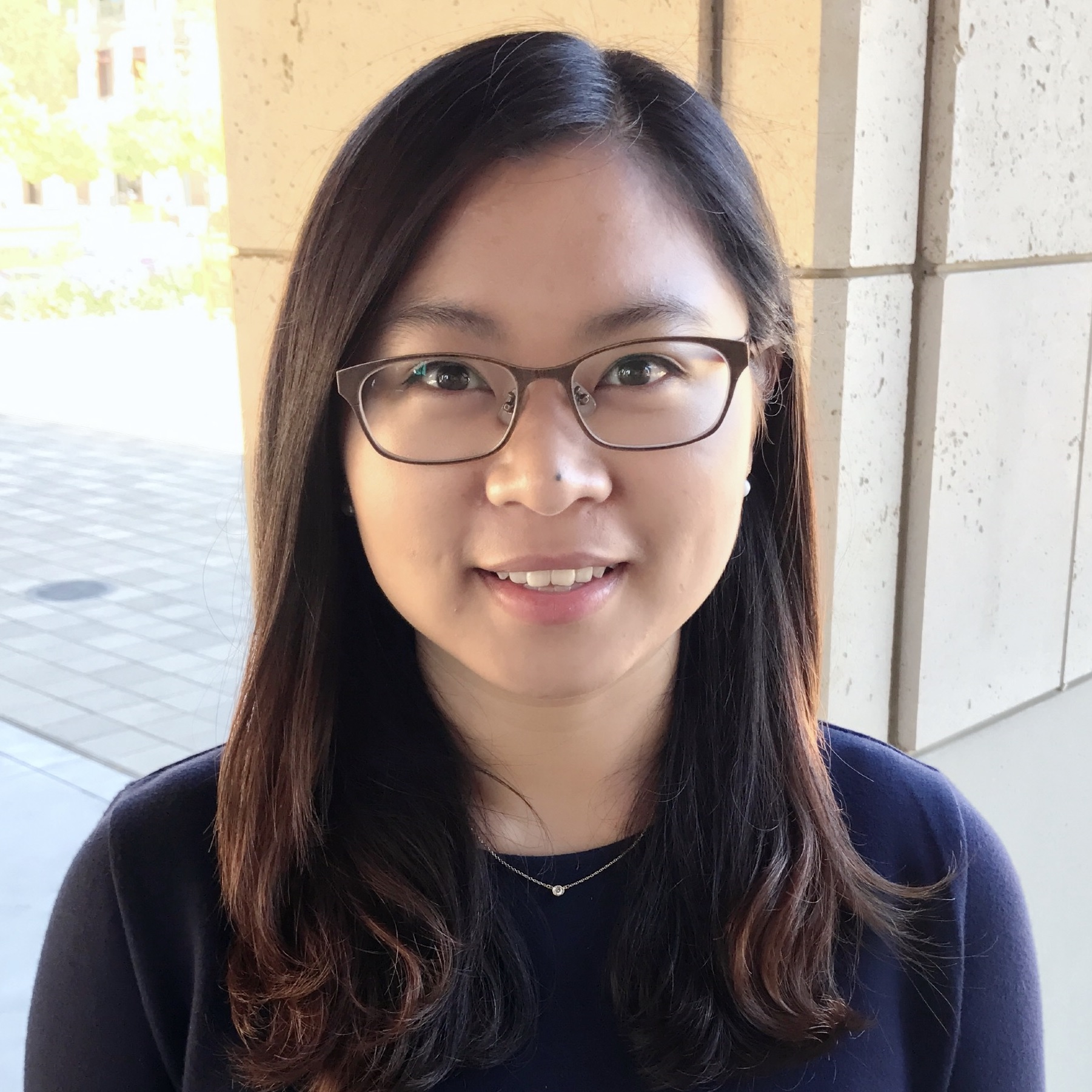 
Thursday, January 18, 3:30pm - 5:00pm, WEL 2.122
Postdoctoral Fellow, Materials Science and Engineering
Stanford University
PhD, Stanford, 2015
Cui Lab: When the size of materials is reduced to the nanoscale dimension, physical and chemical properties can change dramatically. In addition, nanostructures also afford new exciting opportunities of low-cost processing. We are interested in a broad range of nanoscale properties including electronic, photonic, electrochemical, mechanical, catalytic and interfacial properties. Understanding these properties has important technological implications in energy conversion and storage, electronics, biotechnology and environmental technology. We study fundamentals of nanomaterials including nanowires, colloidal nanocrystals and patterned nanostructures, develop low-cost processings and address critical issues in real-world applications.
News Release 8/15/16: SLAC, Stanford Gadget Grabs More Solar Energy to Disinfect Water Faster
h-index: 20 Total Citations: 1252 (Google Scholar Citations, Dec. 2017)
Faculty Recruiting Seminar 
Wednesday, January 10, 3:30pm - 4:30pm, WEL 2.122
Postdoctoral Fellow, Center for Bio-Inspired Energy Research
Northwestern University
ORCID: orcid.org/0000-0001-7757-8335
PhD, Weizmann Institute, Israel, 2013
My current work focuses on ratchets – far-from-equilibrium devices that transport particles using local asymmetries, rather than overall biases. Ratchets are rectifiers – they extract directional motion from non-directed sources of energy, like chemical energy and Brownian motion. Biological motors in the body use ratchet mechanisms, and produce motion very efficiently, even in the highly-damped biological conditions, where the noise is actually orders of magnitude stronger than the chemical energy available. We want to understand how the ratcheting applies to electrons, especially under highly-damped conditions, like in low-mobility organic semiconductors. Very little experimental work has been done on electron ratchets, and so we mainly seek to improve our understanding of the mechanism, with an eye toward possible future applications in solar cells or other electronic devices.
h-index: 5 Total Publications: 14 Total Citations: 165 (ResearcherID, Nov. 2017)
h-index: 5 Total Citations: 212 (Google Scholar Citations, Nov. 2017)
The Seminars page is brought to you by the University Libraries. Our intent is to provide a quick profile of our guest speakers, links to their research group sites, recent publications, author metrics, and other information to enhance your engagement with the guests.
This work is licensed under a Creative Commons Attribution-NonCommercial 4.0 Generic License.